This year, we published a review of studies on the Hybrid Mouse Diversity Panel (HMDP) dataset, a project led by Aldons J. Lusis (David Geffen School of Medicine at UCLA). Our paper in Journal of Lipid Research describes the dataset, summarizes current discoveries facilitated by the dataset, and explains how researchers can use correlation, genetic mapping, and statistical modeling methods with HMDP data to address cardiometabolic questions.
The Hybrid Mouse Diversity Panel (HMDP) is a collection of approximately 100 well-characterized inbred strains of mice that can be used to analyze the genetic and environmental factors underlying complex traits. While not nearly as powerful for mapping genetic loci contributing to the traits as human genome-wide association studies, it has some important advantages. First, environmental factors can be controlled. Second, relevant tissues are accessible for global molecular phenotyping. Finally, because inbred strains are renewable, results from separate studies can be integrated.
Since its development in 2010, studies using the HMDP have validated over a dozen novel genes underlying complex traits. High-throughput technologies have been used to examine the genomes, epigenomes, transcriptomes, proteomes, metabolomes, and microbiomes of mice subjected to various environmental conditions. These analyses have identified many novel genes and significant loci associated with disease risk relevant to obesity, diabetes, atherosclerosis, osteoporosis, heart failure, immune regulation, and fatty liver disease.
The HMDP has substantial potential to advance interdisciplinary research on genetics and computational biology. In order to make HMDP and associated methods accessible to cardiometabolic researchers, our paper includes a glossary of genetics terms and an outline of how the database can be interrogated to address certain questions using correlation, genetic mapping, and statistical modeling.
All of the published data are available and can be readily used to formulate hypotheses about genes, pathways, and interactions. For more information about HMDP, read our article: https://www.ncbi.nlm.nih.gov/pubmed/27099397
The full citation to our paper is:
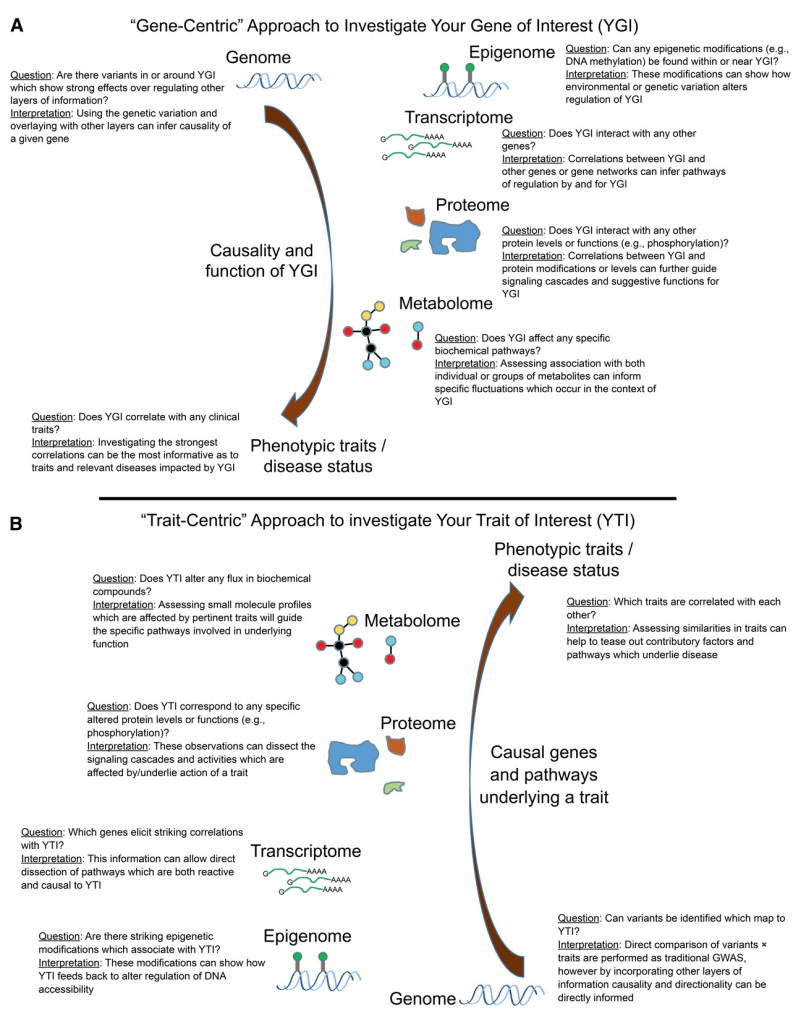
Hypothetical examples of how information from the HMDP can be utilized to explore relationships between genes (A) and traits (B) of interest. Read our paper for more information on methods for exploring their relationships with multiple layers of information.

